Pan-genome analyses of bacteria and archaea are routinely carried out to interpret the within-species variation. However, eukaryotic pan-genome analyses are limited by the large sizes and high complexities of their genomes. Therefore we develloped a new "map-to-pan" strategy ---- EUPAN, specific for eukaryotic pan-genome analysis. EUPAN was primarily developed in the 3000 Rice Genomes Project, in which, EUPAN enables accurate detection of gene presence-absence variations (gene PAVs) for 453 rice accessions at sequencing depth of ~20x. EUPAN suits for pan-genome analysis involving 10 to thousands of individuals from the same or closely related species with medium (~20X) sequencing depth. Moreover, EUPAN can be directly applied to some of current re-sequencing projects primarily aiming to explore single nucleotide variations (SNVs).
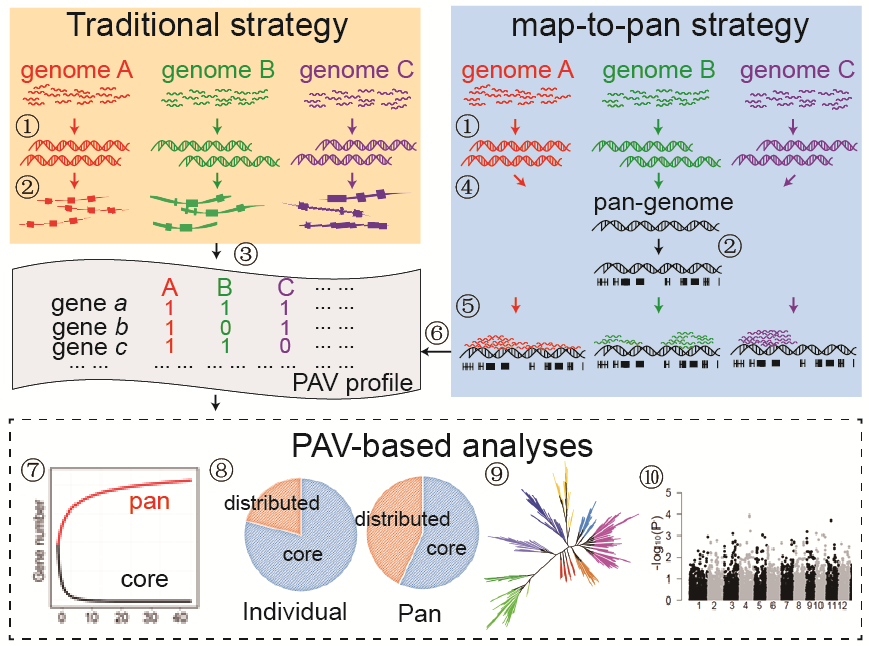
We proposed EUPAN strategy primarily in the 3000 Rice Genomes Project. EUPAN utilizes a "map-to-pan" strategy to determine gene PAVs of each individual genome. EUPAN strategy involves
- the parallel quality control of raw sequencing data;
- de novo assembly of individual genomes;
- construction of pan-genome sequences based on the de novo assemblies and available reference genomes;
- gene annotation of the pan-genome sequences;
- determination of PAVs based on mapping individual reads to pan-genome sequences;
- PAV-based pan-genome analysis.
Eukaryotic large genome studies always involve big data and various softwares; and require very careful parameter selection process. Therfore EUPAN toolbox provides 3 types of tools: 1) single machine version, 2) LSF version (working on supercomputer based on LSF system, in which, "bsub" is used to submit jobs) and 3) SLURM version (working on supercomputer based on SLURM system, in which, "sbatch" is used to submit jobs).
Use & Citation
EUPAN is free for non-commercial use (CC BY-NC 4.0). For commercial use, please contact the authors.
Hu, Z., Sun, C., Lu, K., Chu, X., Zhao, Y., Lu, J., Shi, J., Wei, C., " EUPAN enables pan-genome studies of a large number of eukaryotic genomes", Bioinformatics ,33(15)2408-2409 (2017).
-
Duan, Z., Qiao, Y., Lu, J. et al. HUPAN: a pan-genome analysis pipeline for human genomes. Genome Biol 20, 149 (2019).
Contact Information
Chaochun Wei: ccwei@sjtu.edu.cnNews
- 2017.10.23 version 0.44 released.
- "geneCov" tool now handles more flexible GTF/GFF files.
- 2017.3.1 version 0.43 released.
- Automatic completion with [TAB] for eupan commands.
- 2017.2.22 version 0.42 released.
- Fix a bug in "geneCov" tool.
- Remove redundant plot outputs.
- 2016.10.12 version 0.41 released.
- Fix a bug in "trim" tool.
- 2016.7.20 version 0.4 released.
- Now EUPAN supports SLURM system.
- 2016.4.6 version 0.3 released.
- Add a novel tool ----"sim", to simulate the pan-genome and core-genome size.
- The "geneCov" tool is optimized to generate no intermediate files and it is now faster.
- 2016.3.25 version 0.2 released.
- Fix bugs in the parallelization of the "sam2bam" tool.
- 2016.1.25 version 0.1 released.